How does one Seattle coronavirus patient turn into two six weeks later?
And why does that mean there are hundreds if not thousands of carriers?
How many are infected?
That’s the question many are asking since news broke that a second person in the Seattle area had tested positive for SARS-CoV-2, the novel coronavirus. But along with news of this second confirmed case, we heard that there are likely a few hundred people infected. How could two confirmed cases suggest that there are hundreds infected? The second patient had no history of travel to China, where exposure could have happened. If exposure happened in the Seattle area, was the virus circulating here for a while, infecting not only this person, but many others? Modern genomics tools are providing clues to a mystery. Where did patient 2 catch coronavirus?
Sequencing entire genomes allows us to track randomly occurring mutations in a pathogen — the SARS-CoV-2 virus, in the current situation. Each time a virus reproduces, it copies its genome. Copying a genome is like copying a book by hand – mistakes are bound to be made. Over time, as the virus spreads, tell-tale random mutations pile up at relatively predictable rates. When a mutation is shared between multiple viruses, they likely have a similar source, and the number of mutations that they differ by allows us to estimate the amount of time elapsed between separate infections — a molecular clock.
The clue to solving the Seattle mystery came from the genetic sequencing of SARS-CoV-2 virus from the first two patients to exhibit symptoms in the area. The first patient, who we'll call "patient 1," had returned from Wuhan, China, the epicenter of the global epidemic, on January 15, 2020. This person developed symptoms and was confirmed to be infected with SARS-CoV-2 on January 21. They were isolated at home and advised to minimize contact with others. As it turns out, the first ten confirmed cases in the US had contact with 445 people after symptom onset and before isolation – patient 1 had contact with at least one person, though exactly how many is unclear. Though the CDC regularly checked in with these exposed individuals for 14 days in order to isolate anyone who developed symptoms, it is possible that those who were infected, but developed only very mild or no symptoms, were not tested or isolated, and continued to spread infection in the Seattle area.
The second confirmed infection in the Seattle area was in a high school student. He reported having flu-like symptoms on February 24, six weeks after patient 1, and tested negative for flu at a clinic. His samples were shared with an ongoing research study in the area — the Seattle Flu Study. Remarkably, the test was positive for SARS-CoV-2. Let’s call this person “patient 2.”
Patient 2 had not traveled to China in the past few weeks. Until February 27, you could only get tested for SARS-CoV-2 in the US if you had flu-like symptoms and had returned from China in the past 14 days. The results from the Seattle Flu Study were an important break for US public health — they showed that at least one person with no history of travel to China had been infected with the virus. Where did patient 2 contract the virus? Well, possibly from someone in the community, but we had clearly missed testing and diagnosing that person.

These two cases live in Snohomish County, which borders Seattle's King County. The proximity in space is a clue that the two confirmed SARS-CoV-2 cases may be epidemiologically linked. But they had no known contact. So did the first case, in fact, lead to the second? The genomes of the viruses from patients 1 and 2 were sequenced so researchers could examine mutations in their genomes and link them to each other or to viruses causing outbreaks elsewhere in the world. Remember, these are random mutations, not necessarily leading to any change in the way the virus behaves, but providing important clues about chains of infection. It turned out that the two virus sequences differed by three mutations, but shared one mutation that distinguished them from the genome sequences of viruses isolated from the very first cases in Wuhan in early January 2020.
That shared mutation is a smoking gun. It links patient 1, who returned from Wuhan to Seattle, not only to cases in China but also to patient 2. That one shared mutation tells us that the virus that caused disease in patient 2, could be traced back to the virus that caused disease in patient 1. The shared mutation does not tell us that the two patients were in contact, only that there is a direct line, with an unknown number of people, between the two detected infections. The three mutations between the viruses from patient 1 and 2 matched researchers' expectations exactly for a virus that had been circulating for 6 weeks.
Trevor Bedford, the researcher behind the genome analysis, detailed the work in a blog post, and is quick to point out that the one mutation linking the two cases could have been a chance occurrence. This particular mutation could have been reintroduced into the Seattle area from China. What’s the probability of this having happened? Not nil, it turns out, but a fairly low probability. And because the two confirmed cases in the Seattle area lived close to each other we have greater confidence that they are, in fact, epidemiologically linked.
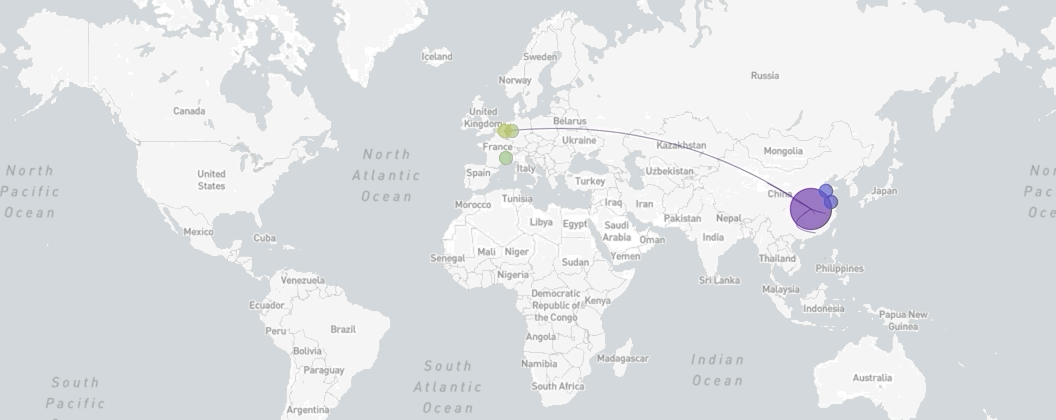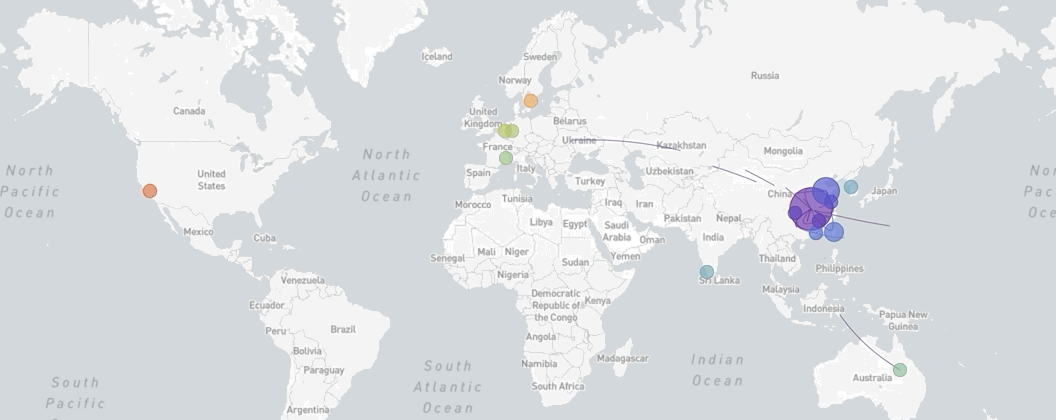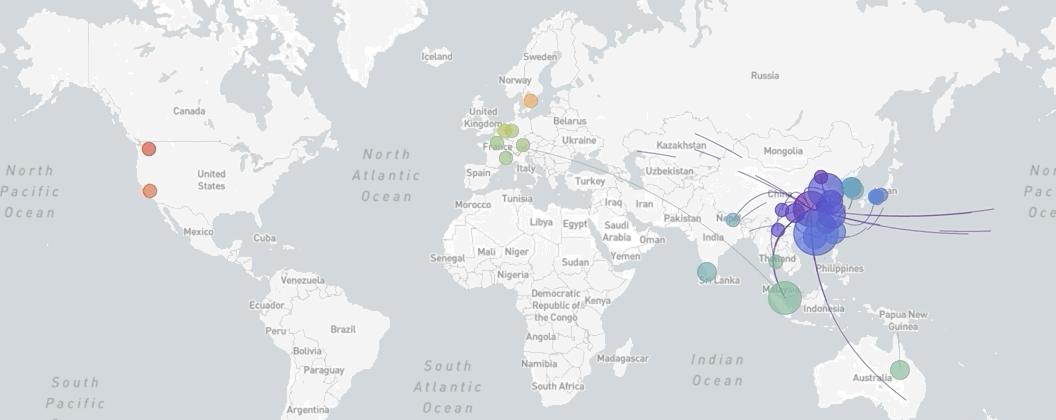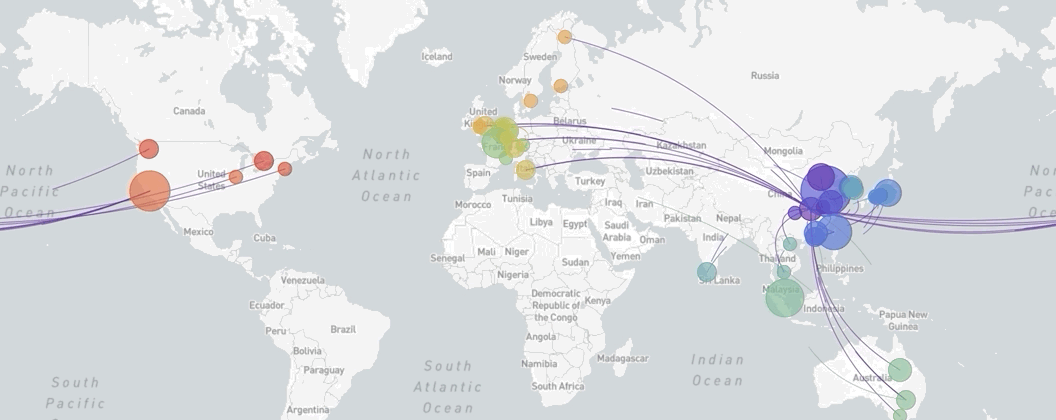
A map of the world showing the spread of SARS-CoV-2, coronavirus, from December 18th, 2019, to March 8th, 2020
Made via Nextstrain
How many people are infected now? Since the diagnosis of patient 2, SARS-CoV-2 viruses from multiple cases in the Seattle area have shared the single mutation found in the first two patients, increasing researchers' confidence that they are linked. If patient 1 transmitted virus to one or more people resulting in an infection in patient 2 on February 24 — i.e. if the virus was being transmitted for six weeks in the community — researchers estimated between 80 and 1500 cases, but likely around 600 people infected as of March 1. We did not detect these cases largely because the US was still scaling up its capacity to test for the virus, and was testing only people with symptoms who had a verifiable history of travel to China in the 14 days prior to symptom onset.
Researchers at the Institute for Disease Modeling in Seattle base this estimate on a simulation that incorporates up-to-date information of how this disease is transmitted from person to person. This includes data being collected and updated by researchers during multiple unfolding epidemics around the world. Data include how many people any given patient infects, the kinds of contact that results in infection, and the time between two successive infections in a chain. From data to-date, SARS-Cov-2 epidemics appear to double in about six days, so 600 cases on March 1st could grow to many thousands of cases in Seattle by month-end in the absence of interventions. The finding of additional virus lineages that may have been circulating in the area would increase the estimated number of people currently infected in Seattle, and hence, the forecasted size of this epidemic.
Human behaviors, including contact mixing (i.e. who interacts with whom) hand washing, and coughing into your elbow, are key to how a respiratory infection epidemic progresses in a population. In a global epidemic, we must also consider differences in human behavior between regions. Do people in Seattle have, on average, more, just as many, or fewer contacts than people in Wuhan? This has implications for how comparable the epidemic in Seattle will be over the next few weeks to the epidemic in Wuhan in January.
Across countries where contact mixing has been studied, people report making, on average, 12-20 contacts each day. Contacts made in households are more often close contact (the kind that facilitates the spread of droplets that viruses can ride on) and these increase with household size. On average, 2.1 people share a household in Seattle, whereas 3 people share a house on average in Wuhan's Hubei province.
According to Jonathan Read, a researcher at Lancaster University who studies disease transmission between people, "Given how SARS-CoV-2 and many other respiratory viruses transmit between people (droplet transmission), it may be the density of your social environment as well as number of the people you encounter that influence transmission risk. The crowding of large families into small households is a good example of this." That said, Read is quick to add "Epidemics are fundamentally driven by human behavior. How people respond to an outbreak, and how they change their behavior in response to public health messages during the course of an epidemic, are possibly more important in determining the course of an epidemic than any underlying ‘peace-time’ differences in social behavior."
China took some very aggressive measures to increase distance between people and reduce viral spread within and outside of Wuhan, including halting air, road, and rail transportation through Wuhan, and closing places where people congregate such as schools, workplaces, and restaurants. This effectively reduced the number of contacts people in Wuhan had largely to only those within households, and along with case finding and contact isolation, slowed the epidemic.
Over the next few weeks we can expect the number infected in Seattle to grow. The King County Public Health Department recommended on March 4th that people older than 60 years of age limit their contact outside the home, and that workplaces enact measures that allow people who can work from home to do so. Amazon, the largest employer in the city, is canceling all non-essential travel for employees and recommending employees work from home. Similarly, Microsoft recommended on March 4th, that its 54,000 Seattle employees work from home for three weeks. Some schools are closed, while parents have been asked to prepare for further closures should children be confirmed with the virus. Human behavior here, as in China, will be driven both by individual capabilities and by societal norms. In China, there appears to have been a prevailing sense of responsibility to contain the disease. To the extent that capabilities and norms differ between the two locations, we can expect the epidemic curve in Seattle to be different from that observed in Wuhan. Bold leadership, strong genomics-informed surveillance, and collective action in Seattle will show if we can slow the epidemic as quickly as Wuhan did.
Supriya Kumar is a public health researcher at the Bill & Melinda Gates Foundation. The comments in this article are her own and do not necessarily reflect the views of the Foundation.

